How Fast Can FEP Run?
by Corin Wagen · Apr 8, 2026

Chariot Race, Jean-Léon Gérôme (1876)
Free-energy-perturbation calculations are notoriously slow. While it's difficult to find public timing benchmarks for most commercial FEP solutions, OpenFE reports speeds of 2–3 hours per RBFE leg, which comports with what we've heard from industry FEP practitioners. (Other sources cite 10 hours per RBFE leg.)
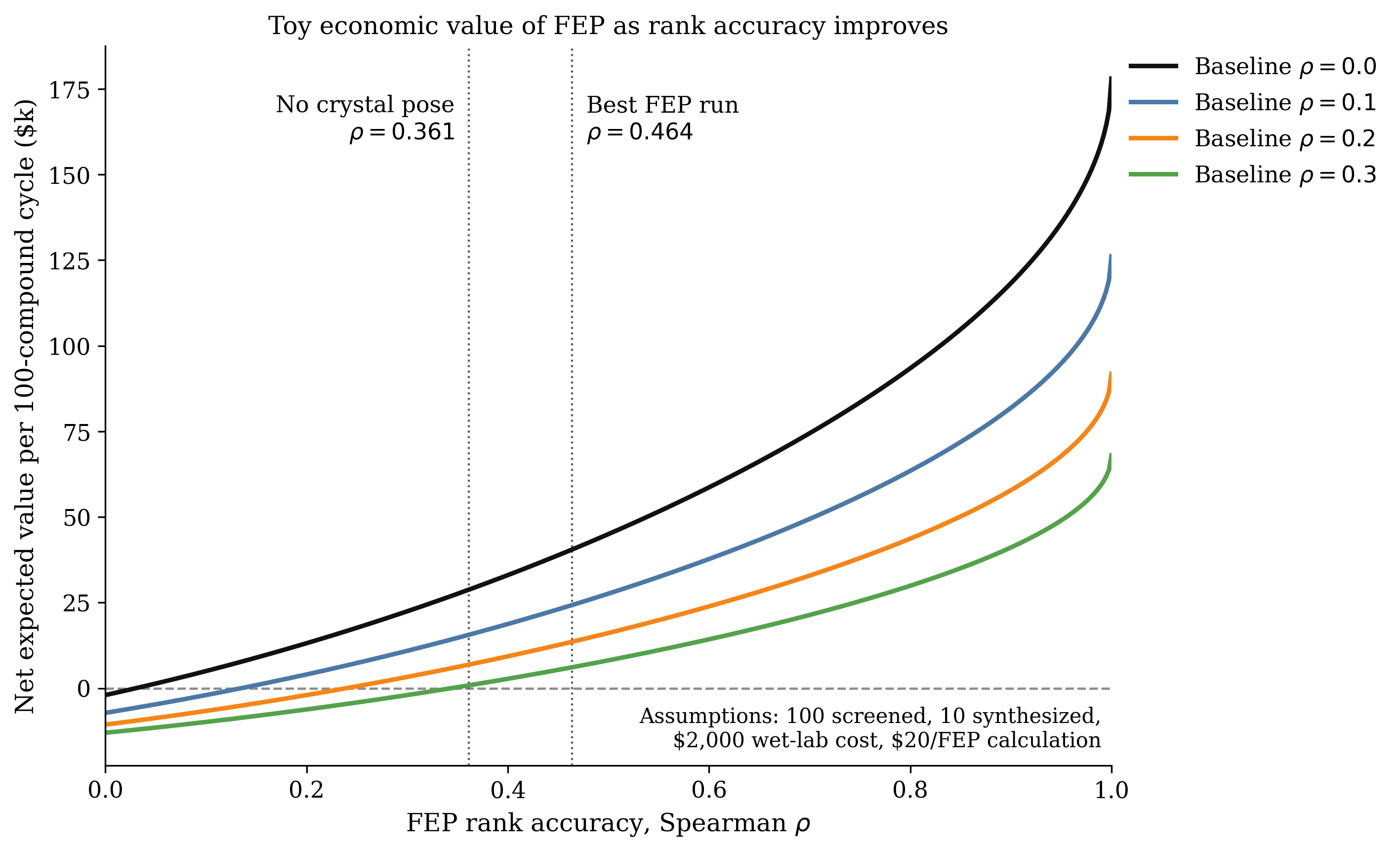
The slow speed of conventional RBFE calculations has important practical implications. Each calculation takes substantial real-world compute resources, which translates into high cost per ligand and limited computational throughput. Accordingly, most drug-discovery teams run relatively few FEP calculations, preferring instead to run large-scale docking, co-folding, or endpoint-based binding-affinity-predicting methods like MM/GBSA. While these faster methods don't generally predict ∆G quantitatively like FEP does, they're still useful for rank-ordering potential binders and prioritizing compounds for FEP and, ultimately, synthesis.
In our FEP launch a few weeks ago, we highlighted how the open-source TMD RBFE engine allowed Rowan to run FEP calculations significantly faster than standard approaches—typically 10–20 minutes per ligand, depending on the system—without compromising accuracy. Here, we want to explore a different question: if we're okay with sacrificing a little accuracy and physical rigor, how fast can Rowan FEP run?
For this blog post, we decided to look at the MCL1 system from the JACS benchmark set. In our original pre-launch FEP testing, MCL1 took 8:31:32 to run on our standard 4x NVIDIA L40S machine, giving a final speed of 12.2 minutes per ligand. With the speed updates we've pushed over the last month, we were able to get the runtime down to 6:00:51 (8.6 minutes per ligand). Here's a public link to the calculation on Rowan.
The above timings are all run with standard settings. Rowan FEP also comes with suggested "fast" settings, which modify the FEP run in several ways:
- The equilibration time for each leg is reduced from 500 ps to 25 ps.
- The simulation time for each leg is reduced from 2 ns to 1 ns.
- The initial number of lambda windows is reduced from 48 to 24 and the minimum overlap/target overlap between lambda windows is reduced from 0.667 to 0.200. (In practice, these changes seem to roughly halve the number of lambda windows used for most transformations.)
If we run MCL1 with these settings, the simulation completes in 01:30:10, or only 2.1 minutes per ligand. Since the entire RBFE graph comprises 79 complex legs and 79 solvent legs, this means that TMD is able to run almost two alchemical legs per minute (158 legs in 90 minutes). Not bad! Here's a link to the calculation on Rowan.
The accuracy of the "fast" FEP run is noticeably worse: cycle-closure errors are higher (from 0.95 kcal/mol to 2.15 kcal/mol), per-leg uncertainties increase, and the overall MAE & RMSE are worse. Fortunately, though, the overall ranking of compounds versus experiment is roughly the same, as assessed by Kendall's τ and Spearman ρ analysis. Visual inspection shows that the "fast" settings exaggerate the slope of the series, under-predicting the affinity of weak binders and over-predicting the affinity of strong binders.
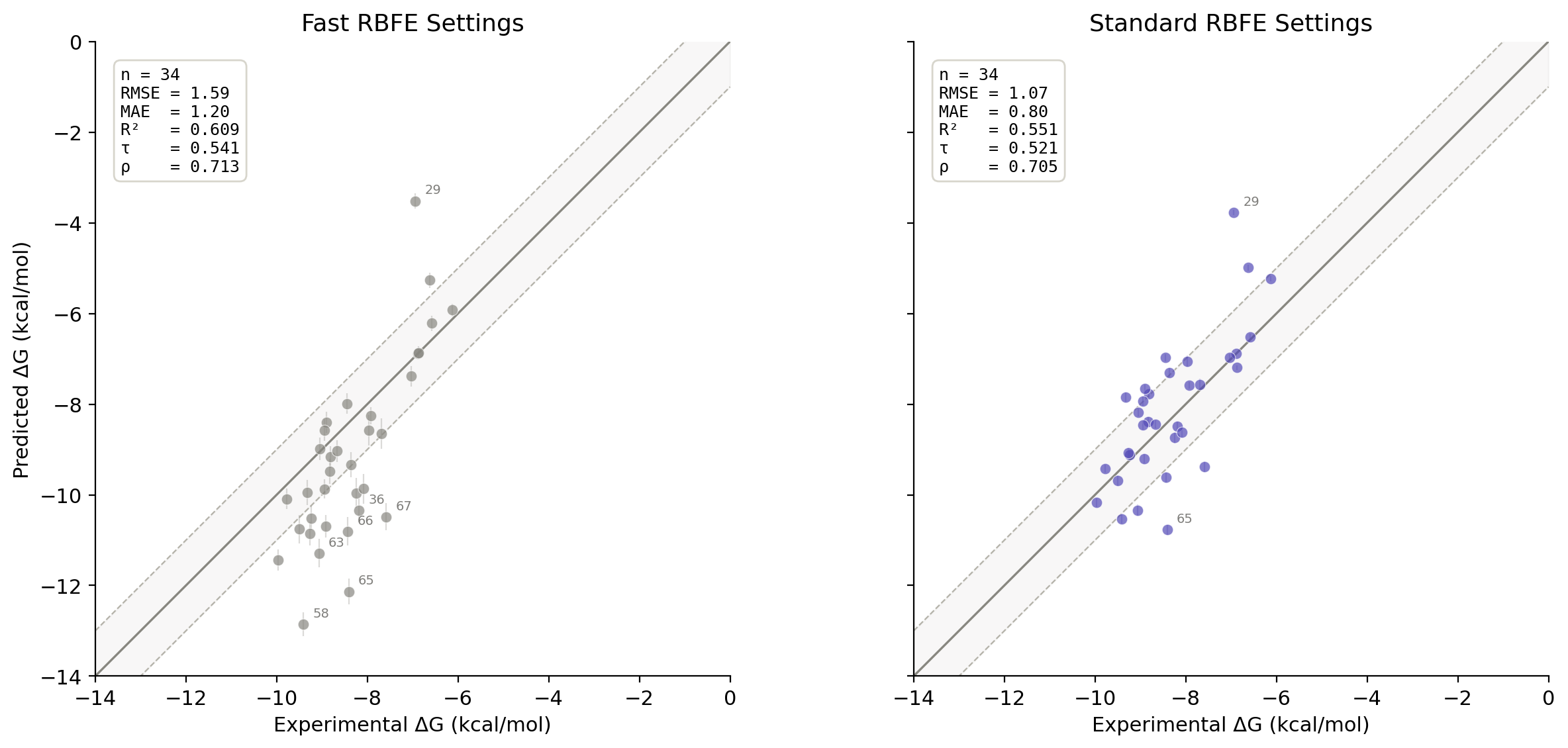
Comparisons of the "fast" and "standard" compounds, excluding the ring-flip compounds.
What can we learn from this brief exercise? Firstly, there's no free lunch here. We can indeed increase the speed of FEP simulations by relaxing how the simulations are run, but we can only do so by introducing increased errors and decreasing the quantitative agreement with experiment. If what we care about is getting our errors as low as possible, this approach doesn't have much to recommend it.
If we're interested in ranking compounds as quickly as possible, though, then we're in luck. At least for this MCL1 test case, running Rowan FEP with speed-optimized settings lets us run simulations approximately 4x faster with minimal loss in rank-based accuracy. And while the final energy error isn't great by FEP standards, it's excellent by the standard of other fast methods like MM/GBSA. (If maximum accuracy is needed, one can simply run "fast" FEP on the entire set of compounds and then re-run the top quartile with standard settings. In this case, this would provide an overall speedup of roughly 2x.)
Further speed improvements could be envisioned via alternate approaches: low-level optimization, fewer RBFE legs, newer GPUs, or active-learning strategies to explore larger compound spaces. We're optimistic that we can increase the speed of Rowan FEP even more in the future, and we're looking forward to the day where FEP can be faster than co-folding.
The above results suggest that "fast" FEP can already excel at rapid ranking of compounds along a series, and we're excited to explore this capability at scale on real project data. If you're interested in bringing Rowan FEP to your drug-discovery project, reach out! We'd love to talk.
Thanks to Forrest York for helpful discussions and for developing the TMD engine.